Overview
Sjögren’s disease is an auto-immune disease affecting 0.1–3% of the population, with women twenty times more likely to develop symptoms such as dry eyes and dry mouth1. The disease is highly heterogeneous, with patients also presenting with a wide range of extraglandular symptoms, and Sjögren’s syndrome is often associated with other auto-immune diseases.
There is a need to better stratify Sjögren’s patients into more clinically relevant subgroups, to develop more personalized and efficacious treatments,
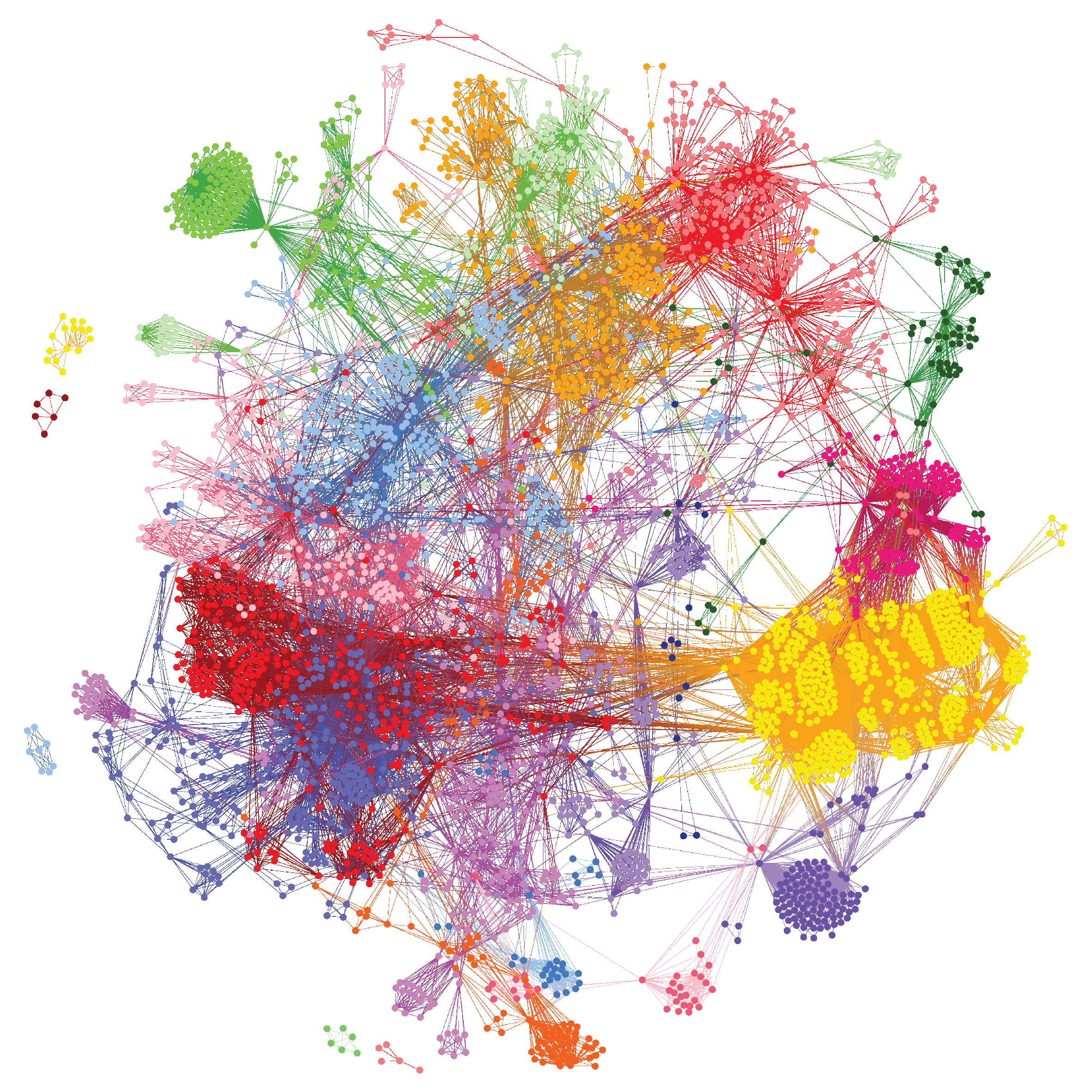
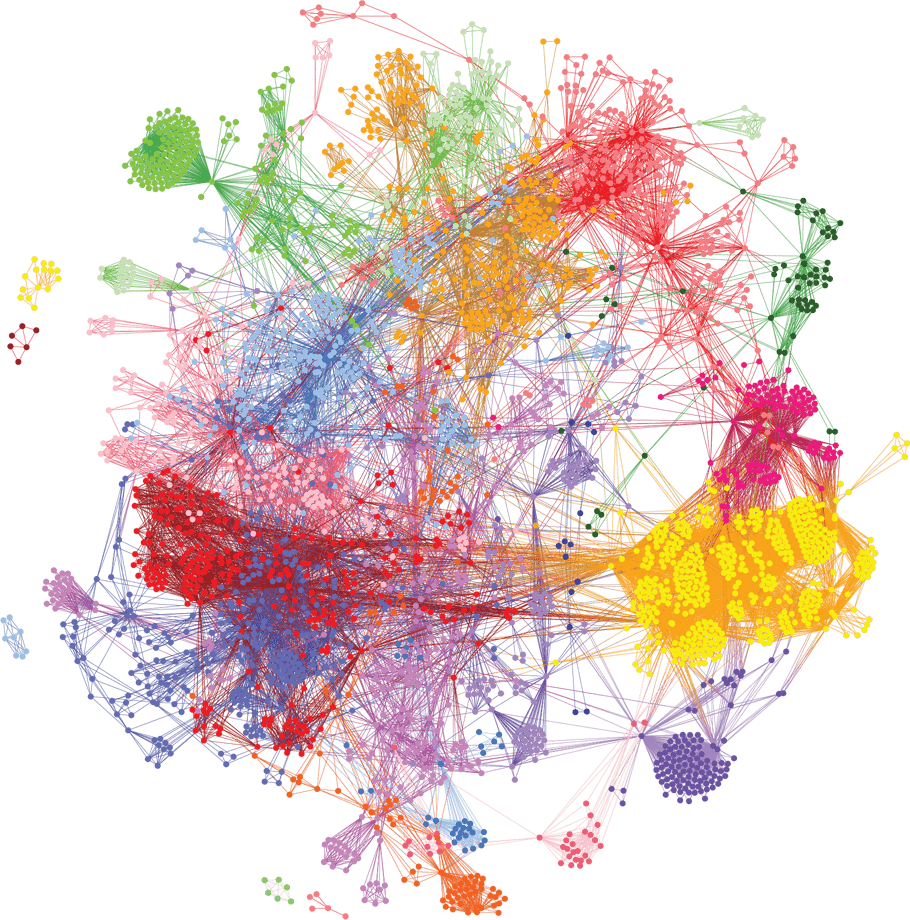
We analyzed genotype data from 990 Sjögren’s cases2, and identified disease-associated combinations of single nucleotide polymorphisms (SNPs). These combinations were then clustered, revealing different patient subgroups that better distinguish and explain the underlying disease mechanisms. We were able to identify gene targets within these subgroups that are involved in a variety of different mechanisms implicated in Sjögren’s disease.