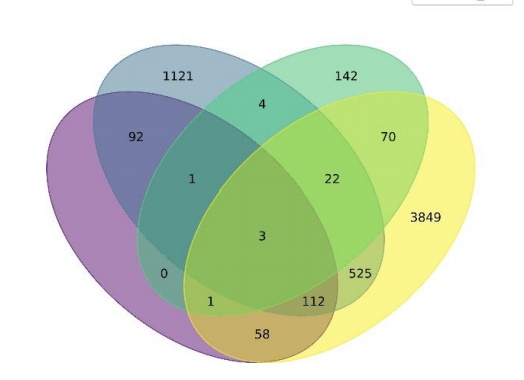
PrecisionLife had the opportunity to share our stratification analysis of Severe Covid-19 Patients with the group.
Fraunhofer Institute for Algorithms and Scientific Computing SCAI,
Bruce Schultz, Christian Ebeling, Vanessa Lage-Rupprecht, Reagon Karki, Yojana Gadiya, Daniel Domingo-Fernández, Manuel Lentzen, Tamara Raschka, Lauren Nicole DeLong, Alpha Tom Kodamullil, Holger Fröhlich, Marc Jacobs & Martin Hofmann-Apitius
ScreeningPort, Fraunhofer Institute for Translational Medicine and Pharmacology ITMP,
Andrea Zaliani, Jeanette Reinshagen, Phil Gribbon & Carsten Claussen in Hamburg and
Gerd Geisslinger & Sandra Ciesek in Frankfurt am Mian
Fraunhofer Cluster of Excellence for Immune Mediated Diseases, CIMD,
Andrea Zaliani, Jeanette Reinshagen, Phil Gribbon, Gerd Geisslinger & Carsten Claussen
Philipp Morris International R&D
Manuel Peitsch & Julia Hoeng
Causality BioModels Pvt Ltd.,
Shounak Baksi
Center for Digital Health, Berlin Institute of Health (BIH), Charité
Sören Lukassen & Roland Eils
Center for Biomedical Data Science, Yale School of Medicine,
Neal G. Ravindra & David van Dijk
Pharmazentrum Frankfurt/ZAFES, Institut Für Klinische Pharmakologie, Klinikum Der Goethe-Universität
Gerd Geisslinger
Institute for Medical Virology, University Hospital
Denisa Bojkova, Jindrich Cinatl & Sandra Ciesek
DZIF, German Centre for Infection Research,
Sandra Ciesek